Two Channel-like Pseudo-Polymorphys Compose of 2,2ÃÆâÃâââ¬Ãâââ¢-Dithiosalicylic Acid and Trialkylammonium
Yang YX, Zhang LI, Li LH, Wen J and Ding KY
Yang YX*, Zhang LI, Li LH, Wen J and Ding KY
Northwest Normal University, College of Chemistry and Chemical Engineering, China
- *Corresponding Author:
- Yang YX
Northwest Normal University, College of Chemistry and Chemical Engineering, China.
Tel: +13893347432
E-mail: yangyx80@163.com
Received Date: November 08, 2016; Accepted Date: December 09, 2016; Published Date: December 15, 2016
Citation: Yang YX, Zhang LI, Li LH, et al. Two Channel-like Pseudo-Polymorphys Compose of 2,2’-Dithiosalicylic Acid and Trialkylammonium. Struct Chem Crystallogr Commun. 2016, 2:2. doi: 10.21767/2470-9905.100023
Abstract
Herein we reported two pseudo-polymorphs of 2,2’-dithiosalicylic acid (C14H10O4S2, DTSA) and trialkylamine with the existence of guanidinium[2C14H9O4S2- ·C+(NH2)3·[HN+(CH3)3] (1)[2C14H9O4S2-·C+(NH2)3·[HN+(C2H5)3] (2). In these two crystal structures that have the same space groups of P¯1, cell parameters, cell volumes and the contents of two asymmetric units are very similar. The only difference is the length of the carbon chains in two trialkyl ammonium cations. And the final stacking patterns of two structures are almost identical, in which two DTSAanions and one guanidinium form 3D channel-like composite host lattices with the help of varied hydrogen bonds and the only trialkyl ammonium is contained among the host lattices to yield the stable crystal structures with the existence of N-H…O interaction. Obviously, in the title compounds, these 3D channel-like host lattices composed of DTSA- anions and guanidinium can adjust their cavities to accommodate different guest molecules to generate the corresponding pseudopolymorphs.
Keywords
Inclusion compound; 2,2'-dithiosalicylic acid; Pseudo-polymorph; Hydrogen bond; 2,2'-Dithiosalicylic acid; DTSA
Introduction
Polymorph is an interesting crystallographic phenomenon. According to the definition of polymorphs, during the crystallization, some ions, atoms, or molecules in the crystalline structures are partly or all substituted with other similar particles, but the types of crystal structures and chemical bonds remain unchanged [1]. Only cell parameters and physical properties of crystals will be linear-related to the change of the particles’ displacement quantity. In modern crystallography, polymorphs are mainly judged by the microscopic symmetries and spatial arrangements of different molecules that are characterized by X-ray diffraction. Normally speaking, polymorphs should belong to the same space group and the related contents of different asymmetry units should be positioned on the same equivalent points. Due to the special properties, polymorphs are widely studied in the pharmaceutical field [2] and other research fields [3]. As a typical host molecule we appreciated, 2,2'-dithiosalicylic acid (C14H10O4S2, DTSA, Scheme 1) that is an aromatic multicarboxylic multi-cyclic molecule with ‘L’ configuration, which is often used as the ligand of MOFs [4-10] and its metal derivatives are useful in medical study [10]. The crystal structure of DTSA has been reported before [11]. The results display that DTSA is able to form different hydrogen bonds by using its carboxyl groups and the central bridging S atoms make the two terminal benzene rings rotate freely to some degree to generate various hydrogen-bonded linking modes. Meanwhile, the rigid benzene rings also can adjust their directions to obtain different stacking patterns and packing modes. Searching in CSD database [12], DTSA can interact with many compounds to form various co crystals, in which DTSA molecules show ‘L’ configuration along the bridging S-S axis and the interplanar angles of two benzene rings are nearly 90°. Additionally, there exist S←O hypervalent bonds between the bridging S atoms and the neighbouring carboxyl O atoms in DTSA to stabilize the ‘L’ type configuration of DTSA [13]. Obviously, as a rigid aromatic multi-cyclic molecule with some flexibility, DTSA can utilize its functional groups to obtain strong hydrogen bonds to act as a potential host molecule with the existence of appropriate guest molecules to get some new inclusion compounds we expected. On the basis of this point, we synthesized two inclusion compound of DTSA and trialkyl amine with the existence of guanidinium, [2C14H9O4S2- ·C+(NH2)3·[HN+(CH3)3](1)and [2C14H9O4S2-·C+(NH2)3·[HN+(C2H5)3] (2). Analysing the crystal structure, compound 1 and 2 belonging to the same space group have very similar crystal parameters and contents of asymmetric units. The host molecule and the auxiliary molecule both form similar 3D host lattices with rectangle cavities with the similar linkage modes of hydrogen bonds and the guest molecules are both contained in the cavities with the existence of N-H…O interaction. In fact, the title compounds are not polymorphs because there are differences in their molecular configurations, hydrogen-bonded linking modes and so on, but they can be regarded as pseudo-polymorphs that can be compared.

Experimental
Synthesis of inclusion compounds
DTSA (98%, A. R.), trimethylaime (99%) and triethylamine (98%) were commercially available from Alfa Aesar. Guanidine hydrochloride was from Tianjin Guangfu Fine Chemical Reagent Institute. DTSA, guanidine hydrochloride and the corresponding amines were dissolved with a 1: 2: 2 molar ratio. The mixture was stirred for about half an hour and set aside to crystallize, finally yielding yellow block crystals of compound 1 and 2 suitable for single crystal X-ray diffraction after about 15 days.
X-ray data collection and structure determination
Crystals of the title compounds were mounted on glass fibers for intensity data collection at room temperature with a Bruker SMART Apex II equipped with a CCD area detector [14]. The structures were solved with the direct methods and refined by full matrix least square methods based on F2, using the structure determination and graphics package SHELXTL [15]. All nonhydrogen atoms were refined with anisotropic displacement parameters, and all H atoms bonded to C atoms were refined at calculated positions, periodic ring on the parent atom. The H atoms bonded to O atoms were located in the difference map and refined. The H atoms bonded to N atoms were firstly searched in the difference map and then refined with the riding model. The crystal and refinement data are given in Table 1. Selected bond lengths and bond angles are listed in Tables 2 and 3.
| Compound | Compound 1 | Compound 2 |
|---|---|---|
| Formula | 2C14H9O4S2-·C+(NH2)3·[HN+(CH3)3 | 2C14H9O4S2-·C+(NH2)3·[HN+(C2H5)3 |
| CCDC No. | ||
| Crystal color | yellow | yellow |
| Crystal shape | block | block |
| Crystal system | Triclinic | Triclinic |
| Space group | P¯ 1 | P¯ 1 |
| Crystal size/mm | 0.30×0.30×0.20 | 0.36×0.23×0.20 |
| a/Å | 10.6646(4) | 10.92620(10) |
| b/Å | 11.5009(4) | 11.69220(10) |
| c/Å | 15.7338(6) | 16.0954(2) |
| α/° | 74.026(2) | 72.0940(10) |
| β/° | 77.683(2) | 78.0580(10) |
| γ/° | 86.228(2) | 82.1410(10) |
| V/Å3 | 1812.52(12) | 1908.35(3) |
| Z | 2 | 2 |
| Dc/(mg·cm-3) | 1.339 | 1.345 |
| µ/mm-1 | 0.315 | 0.303 |
| θ range for data collection | 1.84- 27.68 | 2.44-27.66 |
| Reflection number | 34585 | 11609 |
| Independent reflections | 8425 | 8693 |
| R1,wR2[I>2s(I)] | 0.0372, 0.1003 | 0.0529, 0.1535 |
| R1,wR2(all data) | 0.0490, 0.1099 | 0.0698, 0.1711 |
| S | 1.035 | 1.005 |
Table 1: Crystallographic data.
| Compound 1 | |||
|---|---|---|---|
| O(1)-C(7) | 1.207(2) | S(1)-S(2) | 2.0452(5) |
| O(2)-C(7) | 1.2645(19) | S(2)-C(8) | 1.7881(15) |
| O(3)-C(14) | 1.2764(18) | S(3)-C(15) | 1.7888(17) |
| O(4)-C(14) | 1.2259(19) | S(3)-S(4) | 2.0449(7) |
| O(5)-C(21) | 1.239(2) | S(4)-C(22) | 1.7938(17) |
| O(6)-C(21) | 1.254(2) | N(1)-C(29) | 1.316(2) |
| O(7)-C(28) | 1.308(2) | N(2)-C(29) | 1.312(2) |
| O(8)-C(28) | 1.206(2) | N(3)-C(29) | 1.314(2) |
| S(1)-C(1) | 1.7870(16) | ||
| C(1)-S(1)-S(2) | 104.18(5) | O(5)-C(21)-O(6) | 123.88(16) |
| C(8)-S(2)-S(1) | 105.26(5) | O(8)-C(28)-O(7) | 124.62(15) |
| C(15)-S(3)-S(4) | 105.05(6) | N(2)-C(29)-N(3) | 120.46(18) |
| C(22)-S(4)-S(3) | 105.09(6) | N(2)-C(29)-N(1) | 118.95(19) |
| O(1)-C(7)-O(2) | 124.49(16) | N(3)-C(29)-N(1) | 120.59(19) |
| O(4)-C(14)-O(3) | 122.29(15) | ||
| Compound 2 | |||
| C(1)-S(1) | 1.79(2) | S(3)-C(15) | 1.79(2) |
| N(1)-C(29) | 1.32(4) | S(3)-S(4) | 2.052(9) |
| 1)-C(7) | 1.21(3) | O(4)-C(14) | 1.28(2) |
| S(1)-S(2) | 2.045(8) | S(4)-C(21) | 1.79(2) |
| N(2)-C(29) | 1.32(4) | O(5)-C(27) | 1.24(3) |
| O(2)-C(7) | 1.27(3) | O(6)-C(27) | 1.25(3) |
| S(2)-C(8) | 1.79(2) | O(7)-C(28) | 1.30(3) |
| N(3)-C(29) | 1.31(4) | O(8)-C(28) | 1.21(3) |
| O(3)-C(14) | 1.22(3) | ||
| C(1)-S(1)-S(2) | 104.4(7) | O(5)-C(27)-O(6) | 125(2) |
| C(8)-S(2)-S(1) | 104.7(7) | O(8)-C(28)-O(7) | 124(2) |
| C(15)-S(3)-S(4) | 104.4(8) | N(3)-C(29)-N(2) | 120(3) |
| C(21)-S(4)-S(3) | 104.9(8) | N(3)-C(29)-N(1) | 120(3) |
| O(1)-C(7)-O(2) | 125(2) | N(2)-C(29)-N(1) | 119(4) |
| O(3)-C(14)-O(4) | 122(2) | ||
Table 2: Selected bond lengths (Å) and bong angles (°).
| Hydrogen bonds | O…O distance (Å) | Hydrogen bonds | O…Odistance (Å) |
|---|---|---|---|
| Compound 1 | |||
| N1-H…O5 | 3.053 | N3-H…O3A | 3.130 |
| N1-H…O6C | 2.879 | N3-H…O8B | 2.889 |
| N2-H…O4 | 2.840 | N4-H…O6H | 2.612 |
| N2-H…S2 | 3.514 | O2…H…O2I | 2.443 |
| N2-H…O5 | 2.878 | O3…H…O3A | 2.458 |
| N3-H…O4 | 3.082 | O7-H…O1J | 2.612 |
| H: x, y+1, z; I: 2-x, -1-y, -1+z; J: x-1, y+1, z | |||
| Compound 2 | |||
| N1-H…O6C | 2.808 | N3-H…O5 | 2.788 |
| N1-H…O8B | 3.126 | N4-H…O6 | 2.640 |
| N2-H…O3 | 3.071 | O2…H…O2E | 2.449 |
| N2-H…O4A | 3.106 | O4…H…O4A | 2.448 |
| N2-H…O8B | 2.994 | O7-H…O1F | 2.592 |
| N3-H…O3 | 2.800 | ||
| E: 2-x, -y, 1-z; F: 1-x, 1+y, z | |||
Table 3: Hydrogen-bonding geometry.
Results and Discussion
Crystal structure description
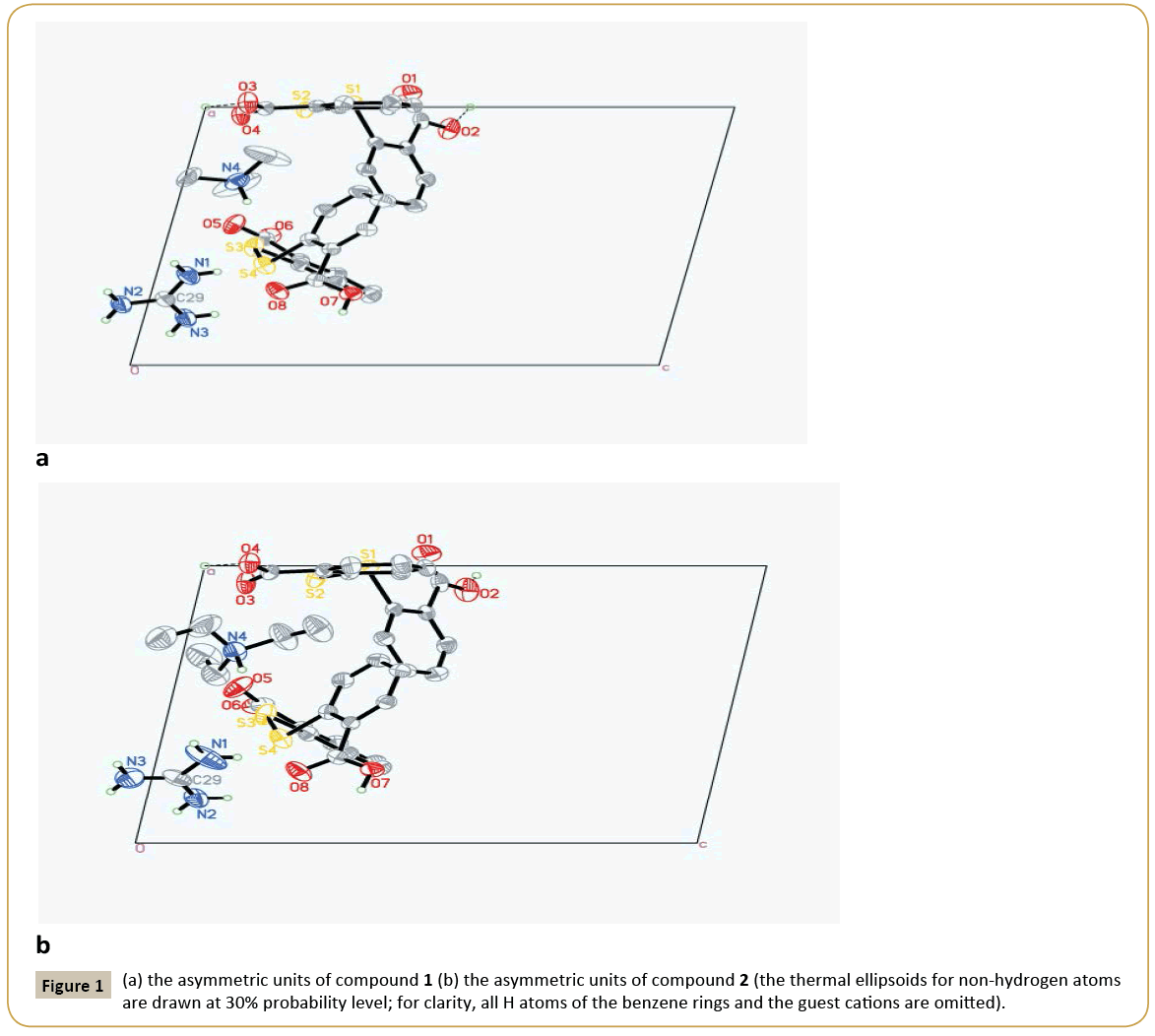
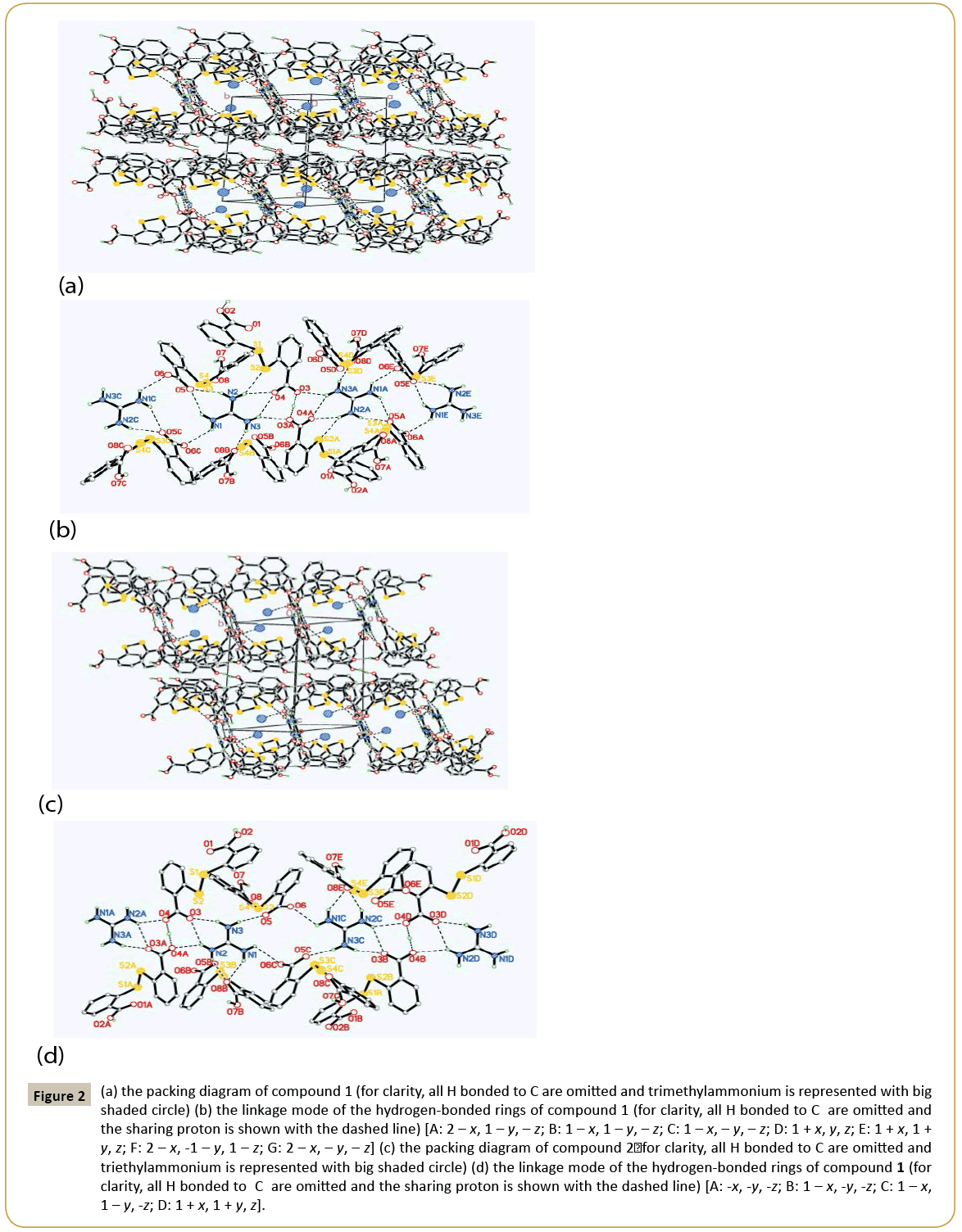
Crystal structure of 1: 2C14H9O4S2-·C+(NH2)3·[HN+(CH3)3]: The asymmetric unit of compound 1 contains two DTSA- anion, one guanidine cation and one trimethylammonium (Figure 1a). Obviously, the N atoms derived from C=N group of guanidine and trimethylammonium accept two protons from DTSA to yield the corresponding cations to counteract the negative charges of DTSA- anion. Observing the structure of the host anion of S1 [the molecule containing S1 atom was designated as S1 and the same representation was adopted below], two benzene rings are coplanar with the mean deviation distances from the leastsquare planes being 0.0029 Å and 0.0079 Å and the interplanar angle if two rings is 79.4°. And two dihedral angles between the carboxyl groups and the related rings are just 4.7° and 9.3° separately, which show the carboxyl groups distort little with the related benzene rings. Notably, the H atoms of the two carboxyl groups in S1 are both located in the special positions. That is to say, the DTSA- anion of S1 shares two protons with another abut symmetry-related anion, which can also be interpreted that S1 anion is a statistic mixture of DTSA molecule and DTSA2- in the crystal structure. And the related C-O bond lengths are varying from 1.207 Å to 1.226 Å and the C=O distances from 1.265 Å to 1.276 Å. At the same time, another anion of S3 displays very similar configuration with S1 , in which the mean deviation values of benzene rings are 0.0056 Å and 0.0059 Å and the dihedral angle of two rings is 82.5°. Interestingly, the torsion angles between the carboxyl groups and the adjacent rings are 4.7° and 30.7°, in which the smaller angle belongs to COO- group with the average C-O distance of 1.239~1.254 Å and the greater one is COOH with the corresponding C-O values of 1.206 Å and 1.308 Å. Clearly, the COOH group retorts more than the COO- group in S3 anion. Additionally, exploring the S←O hypervalent bond lengths (S1←O2: 2.639 Å, S2←O4: 2.675 Å, S3←O5: 2.668 Å, S4←O8: 2.726 Å), it can be found that the weakest hypervalent interaction exists between the C=O motif of COOH group and the neighbouring S atom, which may be attributed to the conjugate structure after the elimination of the proton from the related COOH group. As shown in the packing diagram of compound 1 (Figure 2a), two host anions of DTSA and guanidine cations together form the rectangular channel structure along the direction, in which the anions can be regarded as the ‘top’ and the ‘bottom’ of the channel and the cation as the side faces. Then the two neighbouring channels arranged along the c axis utilize the sharing protons of S1 anions to construct the 3D host lattice and the trimethylammonium is orderly accommodated in the channels to form the stable crystal structure of the inclusion compound. Noticeably, due to the N-H group of the trimethylammonium, there exists strong N-H…O hydrogen bonding between the DTSA- anion and the guest cation, which further strengthens the host-guest interaction to get the steadfast crystal structure. To deeply study the weak interaction of the crystal structure, the hydrogen-bonded linkage mode of the 3D channels is partly shown in Figure 1b. The guanidine cation of N1 forms two pairs of chelated N- H…O hydrogen bonds (N2-H…O4, N3 -H…O4; N2-H… O5, N1 -H…O5) to connect with the anions of S1 and S3, and there also exists one donor hydrogen bond of N-H…S between N2 atom and S2 atom of S1 anion. Meanwhile, the N1 cation also links S3B and S3C with different N-H…O contacts. Based on this, another symmetric center-related N1A cation adopts the same linking pattern to yield the consecutively similar hydrogen-bonded rings as displayed in the Figure 1b. Additionally, the anions of S1 and S1A interact with each other with the sharing proton of O3…H…O3A to further consolidate these continuing hydrogenbonded rings. Then these continuous hydrogen-bonded rings link with each other by a R44(12) ring formed between N1 and N1C. Explicitly, guanidine cation builds the sides of the channel with various hydrogen bonds, while DTSA- anion bonded to guanidine extends widely to construct one half of the roof and bottom of the channel. Equivalently, the host anions of S1 and S3 can connect with other anions to ultimately form the overall channel with the existence of O…H…O interactions (Figure 2b). Therefore, the cooperation of guanidine cation and DTSA- anion finally finishes the architecture of the rectangular channel host lattice shown in the packing diagram. As shown above, guanidine cation serving as the hydrogen bond donor can form several strong hydrogen bonds, and as the side face of the main channel, has a certain rigid plane that can help the host molecules of flexibility to construct channel. In this structure, apart from N-H…O donor hydrogen bond between guanidine cation and DTSA- anion, there also exists a N-H…S hydrogen bond between guanidine and the bridging S atom of DTSA- anion. By calculation, it is also found that the guest trimethylammonium cation can not only form N-H…O hydrogen bond with the host lattice, but also weak C-H…O bonds and C-H…π interactions, which are beneficial to stabilize the crystal structure. In addition, computing with PLATON program, the results elucidate that the potential volume of the cavities is 378.8 Å3 after eliminating the guest cation of trimethylammonium, which occupies 20.9% of the total cell volume.
Figure 2: (a) the packing diagram of compound 1 (for clarity, all H bonded to C are omitted and trimethylammonium is represented with big shaded circle) (b) the linkage mode of the hydrogen-bonded rings of compound 1 (for clarity, all H bonded to C are omitted and the sharing proton is shown with the dashed line) [A: 2 – x, 1 – y, – z; B: 1 – x, 1 – y, – z; C: 1 – x, – y, – z; D: 1 + x, y, z; E: 1 + x, 1 + y, z; F: 2 – x, -1 – y, 1 – z; G: 2 – x, – y, – z] (c) the packing diagram of compound 2ÃÆïÃâüÃâÃâ for clarity, all H bonded to C are omitted and triethylammonium is represented with big shaded circle) (d) the linkage mode of the hydrogen-bonded rings of compound 1 (for clarity, all H bonded to C are omitted and the sharing proton is shown with the dashed line) [A: -x, -y, -z; B: 1 – x, -y, -z; C: 1 – x, 1 – y, -z; D: 1 + x, 1 + y, z].
Crystal structure of 2: 2C14H9O4S2-·C+(NH2)3·[HN+(C2H5)3]: Compound 2 is similar to 1 in the content of the asymmetric unit, and the only difference is the guest cation. Similarly, the angle between two benzene rings in S1 anion is 82.9°, and two dihedral angles between the carboxyl groups and the joint rings are 7.0° and 12.7°, in which one proton is the sharing one and another proton is positional disordered. In S3 anion, the interplanar angle of two rings is 89.0°, and the COO- group distorts 11.0° with the related benzene and the corresponding C-O distances (1.238~1.253 Å) tend to be average after losing the proton, while the COOH group has a greater torsion angle of 23.8° with the related bond lengths being 1.207 Å and 1.298 Å. Moreover, S←O hypervalent bonding interactions in this structure are listed below: S1 ←O1 (2.630 Å), S2←O3 (2.680 Å), S3←O5 (2.644 Å), S1 ←O1 (2.742 Å). Undoubtedly, the carbonyl O of the carboxyl group containing the proton bears the longest S2←O (2.742 Å), which is the same condition as in compound 1. As shown in Figure 2c, it is undoubted that DTSA- anion and guanidine cation in compound 2 form the same channel-like structure as compound 1 and the guest cation of triethylammonium is also contained in the channels with the existence of strong N-H…O interaction to yield the final stable crystal structure. At the same time, it’s also can be seen that the hydrogen bonding linking modes of compounds 2 and 1 are very analogous, but there still are differences such as the absence of N-H…S hydrogen bond, the orientation of the chelated hydrogen bonds of the guanidine cation and the bigger R44(16) hydrogen-bonded ring in compound 2 (Figure 2d). Observing the hydrogen-bonded pattern of compound 2, N3 and N2 atoms of N1 cation produce a pair of chelated hydrogen bonds with O3 atom of S1 to participate the formation of the consecutive hydrogen-bonded rings of N1 and N1A, which has the a like linkage pattern with compound 1. Then N1 and N2 atoms of N1 yield another pair of chelated hydrogen bonds to interact with O8B of S3B to link with the host anion. In succession, N3 and N1 directly take part in the construction of the magnitude ring of R4 4(16) in compound 2, which is distinct with the pattern in compound 1. Similarly, there also exist N-H…O donor hydrogen bond, weak C-H…O hydrogen bond and C-H…π interaction between the guest cation and the host lattice, which undoubtedly contribute to the stability of the final crystal structure. Also, after subtracting the guest cation, the potential cavity volume is 473.1 Å3, which takes up 24.8% of the whole cell volume. It is obvious that the host crystalline lattice can yield larger cavities with the introduction of the larger guest molecule, which illustrates that the same composite host lattice may self-regulate to produce different cavities with varied sizes to accommodate the corresponding guest molecules when introducing various guest templates. Compared the two structures of the title compounds, they both belong to triclinic system with the space group of P¯ 1, and their cell parameters are very similar (compound 1: a=10.6646(4) Å, b=11.5009(4) Å, c=15.7338(6) Å, α=74.026(2)°, β=77.683(2)°, γ=86.228(2)°, V=1812.52(12) Å3, Z=2; compound 2: a=10.92620(10) Å, b=11.69220(10) Å, c=16.0954(2) Å, α=72.0940(10)°, β=78.0580(10)°, γ=82.1410(10)°, V=1908.35(3) Å3, Z=2). Without considering the differences of the guest cations, the molecules in their related cells are located on the almost identical positions (Figure 1). Although the configurations of the host anions are not identical, they both produce very similar hydrogen-bonded patterns during constructing their host lattices, and finally both form the rectangular channels to contain trialkyl ammoniums with different volumes. It is noteworthy that the host anions and guanidine cations together form the channels with negative charges, and it can only contain cations. In addition, the channel can contain two trimethylamine cations, meaning that the channel is large enough and is likely to be able to contain larger cations, such as triethylamine, tetramethyl amine. Clearly, these two structures are not polymorphs strictly according to the definition, but to some degree, they have some characteristics of the polymorphs and we call them pseudopolymorphs for the convenience of comparability. In these two pseudo-polymorphs, DTSA tends to form its related anions under the condition of trialkylamines. The dihedral angles between two benzene rings vary from 79.4° to 89.0° and the torsion angles between the carboxyl groups and the corresponding rings are from 4.7° to 30.7°, which are accordant with the related values reported in the literatures [16-24]. Moreover, S←O hypervalent bond, as an effective three-centre four-electron bond to stabilize DTSA configuration, varies in the range of 2.630 Å~2.762 Å, which is a little more than that of the literature (2.561 Å~2.732 Å) [22]. It is noteworthy that there usually exists strong S←O hypervalent bond between the carbonyl O of the deprotonated carboxyl group and the adjacent S atom. Obviously, there are abundant hydrogen bonds in the title compounds, which are mainly N-H…O hydrogen bonding and O-H…O interactions. In compound 1, the related N…O distances are varying from 2.612 Å to 3.130 Å, and O…O values are between 2.443 Å and 2.612 Å, while the corresponding range of N…O distances is 2.640 Å ~ 3.126 Å and O…O is 2.448 Å ~ 2.592 Å in compound 2. In so many N-H…O hydrogen bonds, the strongest one is formed between the guest cation and the host anion. As to O-H…O interactions, they are stronger than the normal O-H…O hydrogen bonds (O…O distance is 2.6 Å ~ 2.9 Å) in the literature [25]. As mentioned before, in several co crystals of DTSA [11], O-H…N interaction is very common due to the participation of the compounds containing N atoms, and there also exist O-H…O bonding and others in these co crystals, which are not the main driving force in these organic crystals.
Conclusion
To sum up, DTSA, as a L-configuration host molecule with terminal carboxyl groups, can cooperate with suitable auxiliary molecules, such as guanidine cation, to produce hydrogenbonded composite host lattice with 3D cavities. More over the same composite host lattices may form similar cavities with different magnitudes to accommodate the corresponding guest molecules when inducing various guest templates. These results can help us further understand the packing modes of DTSA in the crystal structures and provide the experimental basis for the predesign of the systematic research of the related crystals. The research of other inclusion compounds of DTSA is under way.
Acknowledgements
This work was funded by Young Teacher’s Processing Program of Northwest Normal University (NWNU-LKQN-13-19).
References
- BernsteinJ (2002) Polymorphism in Molecular Crystals. Clarendon, Oxford, UK.
- BrittainHG (2009) Polymorphism in pharmaceutical solids. CRC Press.
- KobayashiY, ItoS, ItaiS, YamamotoK (2000) Physicochemical properties and bioavailability of carbamazepine polymorphs and dehydrate. IntJ Pharm2:137-146.
- Liaw WF, Hsieh CH, Peng SM, Lee GH (2002)SS Bond-activation of diorganyl disulfide by anionic [Mn(CO)5]−: crystal structures of [MnII(SC5H4NO)3]− and [(CO)3Mn(μ-SR)3Co(μ-SR)3Mn(CO)3]− (R=C6H4NHCOPh).InorgChimActa332: 153-159.
- Ramaswamy M, Kanhayalal B, Ganapathi A (2001)Reactions of 2-Mercaptobenzoic Acid with Divalent Alkaline Earth Metal Ions: Synthesis, Spectral Studies, and Single-Crystal X-ray Structures of Calcium, Strontium, and Barium Complexes of2,2’-Dithiobis(benzoic acid). InorgChem27: 6870-6878.
- Lee SM, Wong WT (1996) Oxidative addition of 2,2′-dithiosalicylic acid to the cluster [Os3(CO)10(CH3CN)2]. JCluster Sci7: 37-47.
- Murugavel R, Anantharaman G, Krishnamurthy D, Sathiyendiran M,Walawalkar MG (2000) Extended metal-organic solids based on benzenepolycarboxylic and aminobenzoic acids. Proc Indian Acad Sci Chem Sci 3: 273-290.
- Zhao WN, Zou JW, Yu QS (2004) A one-dimensional ladder-like coordination polymer derived from chains formed via hydrogen bonds: catena-poly[[aquadipyridinenickel(II)]-μ-2,2'-dithiodibenzoato-κ3O,O':O'']. Acta Cryst C60: 443-444.
- Wang ZL, Wei LH, Li MX, Wang JP (2008) Syntheses, Crystal Structures and Theoretical Calculation of Two Transition Metal Dinuclear Complexes. Chinese JStruct Chem11: 1327-1332.
- Henderson W, McCaffrey LJ, Nicholson BK (2000) Synthesis and biological activity of platinum(II) and palladium(II) thiosalicylate complexes with mixed ancillary donor ligands. J Chem SocDalton Trans: 2753-2760.
- Humphrey SM, Wood PT (2003) 2,2'-Disulfanyldibenzoic acid. Acta CrystE9:1364-1366.
- Allen FH, Davies JE, Johnson O, Kennard O(1991) The development of versions 3 and 4 of the Cambridge Structural Database System. J Chem Inf Comput Sci 2: 187-204.
- Broker GA, Tiekink ERT (2007) Co-crystal formation between 2,2′-dithiodibenzoic acid and each of4,4′-bipyridine, trans-1,2-bis(4-pyridyl)ethene and 1,2-bis(4-pyridyl)ethane. CrystEngComm 9: 1096-1109.
- SMART APEX Software (2016) 5.624 for SMART APEX detector. Bruker Axs Inc., Madison, Wisconsin, USA.
- Sheldrick GM (2000) SHELXTL Version 6.10. Bruker AXS Inc., Madison, Wisconson, USA.
- Meng XG, Xiao YL, Zhang H, Zhou CS (2008) Hydrogen-bonded layers in the 1:2 cocrystal of 2,2'-dithiodibenzoic acid with isonicotinohydrazide. Acta Cryst C64:261-263.
- Basiuk EV, LaraJ, BasiukVA, ToscanoR (1999)Complexes of1,4,10,13-tetraoxa-7,16-diazacyclooctadecane (4,13-diaza-18-crown-6) with 2,5-pyridinedicarboxylic and 2,2′-dithiosalicylic acids. J Chem Crystallogr 29: 1157-1163.
- Hu RF, Wen YH, Zhang J, Li ZJ, Yao YG, et al. (2004) 4,4'-Bipyridylium bis(hydrogen 2,2'-dithiodibenzoate) dihydrate. Acta Cryst E60:2029-2031.
- BrokerGA, Tiekink ERT (2007)Co-crystal formation between 2,2′-dithiodibenzoic acid and each of 4,4′-bipyridine, trans-1,2-bis(4-pyridyl)ethene and 1,2-bis(4-pyridyl)ethane. CrystEngComm9: 1096-1109.
- ArmanHD, MillerT, Poplaukhin P, Tiekink ERT (2010) 2,2'-(Disulfanediyl)dibenzoic acid-N,N'-bis(3-pyridylmethyl)ethanediamide (1/1). Acta CrystallogrE66:2590-2591.
- Kresinski RA, Fackler Jr (1994) 2,2'-Dithiodisalicyclic acid tetrahydrofuran solvate. Acta CrystallogrC50: 2039-2041.
- FazilS, RavindranR, Devi AS, Bijili BK (2012) Structural studies of 1-phenyl-2,3-dimethyl-5-oxo-1,2-dihydro-1H-pyrazol-4-ammonium 2[(2-carboxyphenyl) disulfanyl]benzoate.JMolStruct1021: 147-152.
- YangGX, Zhang SZ, Wang ZL (2010)Crystal structure of bis(2-aminopyridinium) 2,2′-dithiobis(benzoate) dihydrate, [C5H7N2]2[C14H8O4S2] · 2H2O.New Crystal Structures225: 367-368.
- Liu Z, Liu Q, Yuan L, Liu W (2010)1,1'-(p-Phenylenedimethylidene)diimidazol-3-ium bis{2-[(2-carboxyphenyl)disulfanyl]benzoate} dihydrate.Acta Crystallogr 66:3230.
- MontbrunD, Le BG, Mason SA, Prange T, Lesieur S,et al.(2013) A highly hydrated α-cyclodextrin/1-undecanol inclusion complex: crystal structure and hydrogen-bond network from high-resolution neutron diffraction at 20 K.Acta Crystallogr B69: 214-227.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences